www.acsprf.org
Reports: DNI350270-DNI3: Molecular Mechanism of Methane to Methanol Conversion in Soluble Methane Monooxygenase
Serena DeBeer, PhD , Cornell University
Biologically, the conversion of methane to methanol is carried out by methane monoxygenases (MMOs) found in methantrophic bacteria. The soluble form of MMO utilizes a binuclear iron active site, which reacts with dioxygen proceeding through a series of intermediates. These include a mu-peroxo-Fe(III)Fe(III) species (MMO-P) and then an Fe(IV)Fe(IV) species (MMO-Q), which is responsible for methane hydroxylation. During the last year, our goal has been to use X-ray spectroscopic methods in order to obtain fundamental mechanistic insights into the enzymatic conversion of methane to methanol. We have focused on the further development of X-ray emission spectroscopy as a means to this end. We have studied both mononuclear and binuclear model complexes and closely correlated our experimental results to theory in order to further the information content of these spectra. We have also initiated studies on the MMO enzyme system itself. The preliminary data for which are also described below.
Studies of mononuclear model complexes by XES. During the current grant period, we have advanced our understanding of XES spectra through detailed experimental and theoretical studies of monoculear Fe complexes. The valence to core region of the spectrum results from valence electrons refilling an Fe 1s core hole, which is created by ionization from a high energy X-ray. The resultant emission processes may be correlated in a simple picture to a one electron molecular orbital picture. Using ferrocene and ferrocenium as "classic" well-defined spectroscopic references, we have demonstrated that these spectra arise from dipole allowed metal 4p character, which mixes into ligand based orbitals. The relative energies of the resultant transitions can be correlated to the energetics of ligand-based molecular orbitals and hence serve as a sensitive probe of ligand identity. These results were published in Inorganic Chemistry earlier this year (K. M. Lancaster, K. D. Finkelstein, S. DeBeer, Inorg. Chem., 2011, 50, 6767).
In addition, we have examined the sensitivity of XES to the nature of the bound ligands (i.e. sigma-donating, pi-donating, and pi-accepting). Using a combination of theoretical metal and ligand XES calculation, we have clearly established the ligand-based origin of valence to core spectral features and shown that the intensity mechanism is dominantly sigma-based in origin. This approach has also been applied to test the potential of XES methods for understanding mononuclear enzymatic intermediates. Specifically, we have investigated the mechanism of extradiol dioxygenase family of enzymes that catalyze the oxidative ring cleavage of catechol derivatives. We have shown that XES should be able to distinguish different intermediates within the reaction cycle. For instance a side-on vs end-on superoxide should be distinguishable due to differing levels of stabilization and Fe 4p mixing for the various orbitals interacting with the metal. The side-on conformation would be expected to interact strongly through the oxygen 2p-2p pi bonding orbital with a weaker interaction between the 2p-2p pi*; the end-on species, in contrast, would bond strongest through 2p-2p pi* orbitals with the 2p-2p pi forming a secondary interaction. The differential stabilization of, and Fe 4p contributions to, these orbitals would lead to valence-to-core XES features with characteristic energies and intensities that could be used to distinguish between the two orientations. Similarly possible lactone and dioxetane intermediates could be distinguished. The results of this work were published in JACS earlier this year (C.J. Pollock and S. DeBeer, JACS, 2011, 133, 5594), and form a foundation for the extension of these methods to both binuclear models and the enzymatic systems described below.
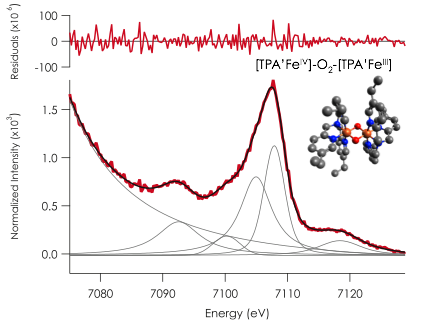
Preliminary Studies of binuclear models and MMO. Our studies of mononuclear iron complexes by XES have been extended to binuclear models prepared in our laboratories (based on literature syntheses from L. Que, Jr.). Figure 1 below shows data on an iron bis-mu-oxo complex, which structurally may bear some similarity to MMO intermediate Q. Interestingly the data show a clear satellite feature at 7092 eV which is attributed to an O 2s to Fe 1s transition, based on parallel correlations with our mononuclear studies. Our preliminary data suggests that if intermediate Q is a bis-mu-oxo type structure, it should be possible to establish this by XES. Our studies on mononuclear systems (described above) suggest alternate O2-binding modes should also be distinguishable.
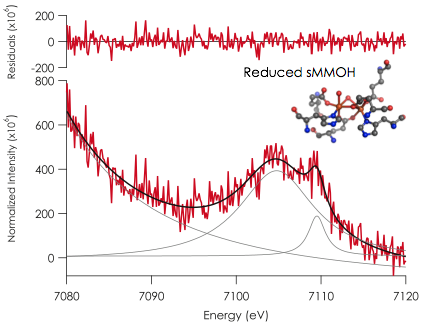
In addition, preliminary data on reduced MMO have been obtained. Due to the low concentration (and in the case of the intermediates, susceptibility to reduce), obtaining data on the protein presents significant challenges. However, our initial results are promising and indicate that with sufficient beam time, insights into the enzymatic intermediates can be achieved utilizing combined experimental and theoretical methods. On going experimental and theoretical work on the enzyme system is in progress in our laboratories.
Figure 1. V2C Kµ XES of a binuclear iron model complex bearing two bridging oxos. Both one iron is LS FeIII and the other is S = 1 FeIV, ferromagnetically coupled to give a net S= 3/2 complex. V2C data represents an average of 12 scans recorded at CHESS. Data were recorded from neat sample of solid compound.
Figure 2. V2C Kµ XES of reduced form of soluble methane monooxygenase hydrolase. Both irons are HS FeII. Data represent the average of 16 scans at SSRL on frozen solution samples containing 3.34 mM Fe. Data statistics are poor, and no Kµ" can be resolved.